Chapter 02: Inference Algorithms#
This notebook explores the different Bayesian inference algorithms available in vangja. We’ll use the multiplicative model from the Air Passengers dataset to compare:
Maximum A Posteriori (MAP) — Point estimates
Variational Inference (ADVI) — Approximate posterior
Markov Chain Monte Carlo (NUTS) — Full posterior sampling
Each method offers different trade-offs between computational speed and the richness of uncertainty quantification.
Reference: For a detailed exploration of these Bayesian inference techniques in time series forecasting, see Krajevski & Tojtovska Ribarski (2026): Going NUTS with ADVI: Exploring various Bayesian Inference techniques with Facebook Prophet, arXiv:2601.20120.
Setup and Imports#
[1]:
import warnings
warnings.filterwarnings("ignore")
[2]:
import time
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from vangja import FourierSeasonality, LinearTrend
from vangja.datasets import load_air_passengers
from vangja.utils import metrics
# Set random seed for reproducibility
np.random.seed(42)
print("Imports successful!")
Imports successful!
Load and Prepare Data#
We’ll use the same Air Passengers dataset from the Getting Started notebook.
[3]:
# Load Air Passengers dataset from vangja.datasets
air_passengers = load_air_passengers()
# Split data: use last 12 months for testing
train = air_passengers[:-12].copy()
test = air_passengers[-12:].copy()
print(
f"Training set: {train['ds'].min()} to {train['ds'].max()} ({len(train)} samples)"
)
print(f"Test set: {test['ds'].min()} to {test['ds'].max()} ({len(test)} samples)")
Training set: 1949-01-01 00:00:00 to 1959-12-01 00:00:00 (132 samples)
Test set: 1960-01-01 00:00:00 to 1960-12-01 00:00:00 (12 samples)
Define the Multiplicative Model#
We’ll use a multiplicative model that captures the increasing variance with trend:
where \(g(t)\) is the linear trend and \(s(t)\) is the combined seasonality.
[4]:
def create_model():
"""Create a fresh multiplicative model instance."""
return LinearTrend(n_changepoints=25) ** (
FourierSeasonality(period=365.25, series_order=10)
+ FourierSeasonality(period=7, series_order=3)
)
print(f"Model structure: {create_model()}")
Model structure: LT(n=25,r=0.8,tm=None) * (1 + FS(p=365.25,n=10,tm=None) + FS(p=7,n=3,tm=None))
1. Maximum A Posteriori (MAP) Estimation#
What is MAP?#
Maximum A Posteriori (MAP) estimation finds the single most probable parameter values given the data and priors. Mathematically, MAP finds:
where \(P(D | \theta)\) is the likelihood and \(P(\theta)\) is the prior.
Advantages:
Very fast computation
Good for quick prototyping and model comparison
Works well when you have enough data
Limitations:
Provides only a point estimate (no uncertainty quantification)
Can get stuck in local optima
Doesn’t capture the full shape of the posterior distribution
PyMC’s map vs pymc-extras’ mapx#
Vangja supports two MAP implementations:
Feature |
|
|
|---|---|---|
Backend |
SciPy optimizers |
JAX-based optimization |
Speed |
Slower |
Significantly faster |
Gradient computation |
Numerical or PyTensor |
JAX autodiff |
Recommended |
Legacy support |
Default choice |
The mapx method from pymc-extras uses JAX for automatic differentiation, resulting in much faster optimization, especially for models with many parameters.
1.1 MAP with mapx (Recommended)#
[5]:
# Create and fit the model using mapx (default)
model_mapx = create_model()
start_time = time.time()
model_mapx.fit(train, method="mapx")
mapx_time = time.time() - start_time
print(f"MAPX fitting time: {mapx_time:.2f} seconds")
WARNING:2026-02-26 20:45:49,639:jax._src.xla_bridge:876: An NVIDIA GPU may be present on this machine, but a CUDA-enabled jaxlib is not installed. Falling back to cpu.
MAPX fitting time: 8.82 seconds
[6]:
# Predict and evaluate
future_mapx = model_mapx.predict(horizon=365, freq="D")
metrics_mapx = metrics(test, future_mapx, "complete")
print("MAPX Metrics:")
display(metrics_mapx)
MAPX Metrics:
| mse | rmse | mae | mape | |
|---|---|---|---|---|
| series | 629.010786 | 25.080087 | 21.254398 | 0.04267 |
1.2 MAP with map (Standard PyMC)#
[7]:
# Create and fit the model using standard PyMC map
model_map = create_model()
start_time = time.time()
model_map.fit(train, method="map")
map_time = time.time() - start_time
print(f"MAP fitting time: {map_time:.2f} seconds")
print(f"Speedup with mapx: {map_time / mapx_time:.1f}x faster")
MAP fitting time: 6.71 seconds
Speedup with mapx: 0.8x faster
[8]:
# Predict and evaluate
future_map = model_map.predict(horizon=365, freq="D")
metrics_map = metrics(test, future_map, "complete")
print("MAP Metrics:")
display(metrics_map)
MAP Metrics:
| mse | rmse | mae | mape | |
|---|---|---|---|---|
| series | 648.633502 | 25.468284 | 21.179644 | 0.042348 |
2. Variational Inference (ADVI)#
What is Variational Inference?#
Variational Inference (VI) approximates the true posterior distribution \(P(\theta | D)\) with a simpler distribution \(Q(\theta)\) from a tractable family. The goal is to minimize the Kullback-Leibler (KL) divergence between \(Q\) and the true posterior:
Automatic Differentiation Variational Inference (ADVI)#
ADVI is a specific VI algorithm that:
Transforms all parameters to unconstrained space
Approximates the posterior with a multivariate Gaussian
Uses gradient-based optimization to find the best approximation
Advantages:
Much faster than MCMC
Provides uncertainty estimates (unlike MAP)
Scales well to large datasets
Limitations:
Approximation may miss multimodal posteriors
Gaussian assumption may be too restrictive
Can underestimate uncertainty
Variants#
Method |
Description |
|---|---|
|
Mean-field approximation (diagonal covariance) |
|
Full covariance matrix (captures correlations) |
[9]:
# Create and fit the model using ADVI
model_advi = create_model()
start_time = time.time()
model_advi.fit(
train,
method="advi",
n=50000, # number of ADVI iterations
samples=1000, # number of posterior samples to draw after ADVI convergence
)
advi_time = time.time() - start_time
print(f"ADVI fitting time: {advi_time:.2f} seconds")
Finished [100%]: Average Loss = -7.8993
ADVI fitting time: 37.00 seconds
[10]:
# Predict and evaluate
future_advi = model_advi.predict(horizon=365, freq="D")
metrics_advi = metrics(test, future_advi, "complete")
print("ADVI Metrics:")
display(metrics_advi)
ADVI Metrics:
| mse | rmse | mae | mape | |
|---|---|---|---|---|
| series | 490.516804 | 22.147614 | 19.328256 | 0.040094 |
3. Markov Chain Monte Carlo (NUTS)#
What is MCMC?#
Markov Chain Monte Carlo (MCMC) methods generate samples from the posterior distribution by constructing a Markov chain that has the posterior as its stationary distribution. After enough iterations, the samples approximate the true posterior.
No-U-Turn Sampler (NUTS)#
NUTS is an advanced MCMC algorithm that:
Uses Hamiltonian dynamics to propose distant moves
Automatically tunes the trajectory length (no manual tuning needed)
Avoids random walk behavior, leading to efficient exploration
Advantages:
Provides the gold standard for posterior inference
Captures the full posterior distribution, including multimodality
Accurate uncertainty quantification
Diagnostic tools available (R-hat, ESS, divergences)
Limitations:
Computationally expensive
Requires tuning (warmup/burn-in period)
May have convergence issues for complex models
Sampler Backends#
Vangja supports multiple NUTS backends:
Backend |
Description |
|---|---|
|
Default PyMC sampler |
|
Fast Rust-based sampler |
|
JAX-based sampler (GPU support) |
|
JAX-based sampler |
[11]:
# Create and fit the model using NUTS
# Using fewer samples for demonstration (increase for production)
model_nuts = create_model()
start_time = time.time()
model_nuts.fit(
train,
method="nuts",
samples=1000, # Number of posterior samples per chain
chains=4, # Number of independent chains
cores=4, # Parallel cores to use
)
nuts_time = time.time() - start_time
print(f"NUTS fitting time: {nuts_time:.2f} seconds")
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [lt_0 - slope, lt_0 - intercept, lt_0 - delta, fs_0 - beta(p=365.25,n=10), fs_1 - beta(p=7,n=3), sigma]
Sampling 4 chains for 1_000 tune and 1_000 draw iterations (4_000 + 4_000 draws total) took 227 seconds.
NUTS fitting time: 230.79 seconds
[12]:
# Predict and evaluate
future_nuts = model_nuts.predict(horizon=365, freq="D")
metrics_nuts = metrics(test, future_nuts, "complete")
print("NUTS Metrics:")
display(metrics_nuts)
NUTS Metrics:
| mse | rmse | mae | mape | |
|---|---|---|---|---|
| series | 698.054001 | 26.420712 | 22.329609 | 0.044714 |
Comparison of Results#
Let’s compare all inference methods side by side.
[13]:
# Combine all metrics
comparison = pd.DataFrame(
{
"Method": ["MAPX", "MAP", "ADVI", "NUTS"],
"Time (s)": [mapx_time, map_time, advi_time, nuts_time],
"RMSE": [
metrics_mapx["rmse"].values[0],
metrics_map["rmse"].values[0],
metrics_advi["rmse"].values[0],
metrics_nuts["rmse"].values[0],
],
"MAE": [
metrics_mapx["mae"].values[0],
metrics_map["mae"].values[0],
metrics_advi["mae"].values[0],
metrics_nuts["mae"].values[0],
],
"MAPE": [
metrics_mapx["mape"].values[0],
metrics_map["mape"].values[0],
metrics_advi["mape"].values[0],
metrics_nuts["mape"].values[0],
],
}
)
print("\nComparison of Inference Methods:")
display(comparison)
Comparison of Inference Methods:
| Method | Time (s) | RMSE | MAE | MAPE | |
|---|---|---|---|---|---|
| 0 | MAPX | 8.824514 | 25.080087 | 21.254398 | 0.042670 |
| 1 | MAP | 6.714782 | 25.468284 | 21.179644 | 0.042348 |
| 2 | ADVI | 37.004420 | 22.147614 | 19.328256 | 0.040094 |
| 3 | NUTS | 230.786814 | 26.420712 | 22.329609 | 0.044714 |
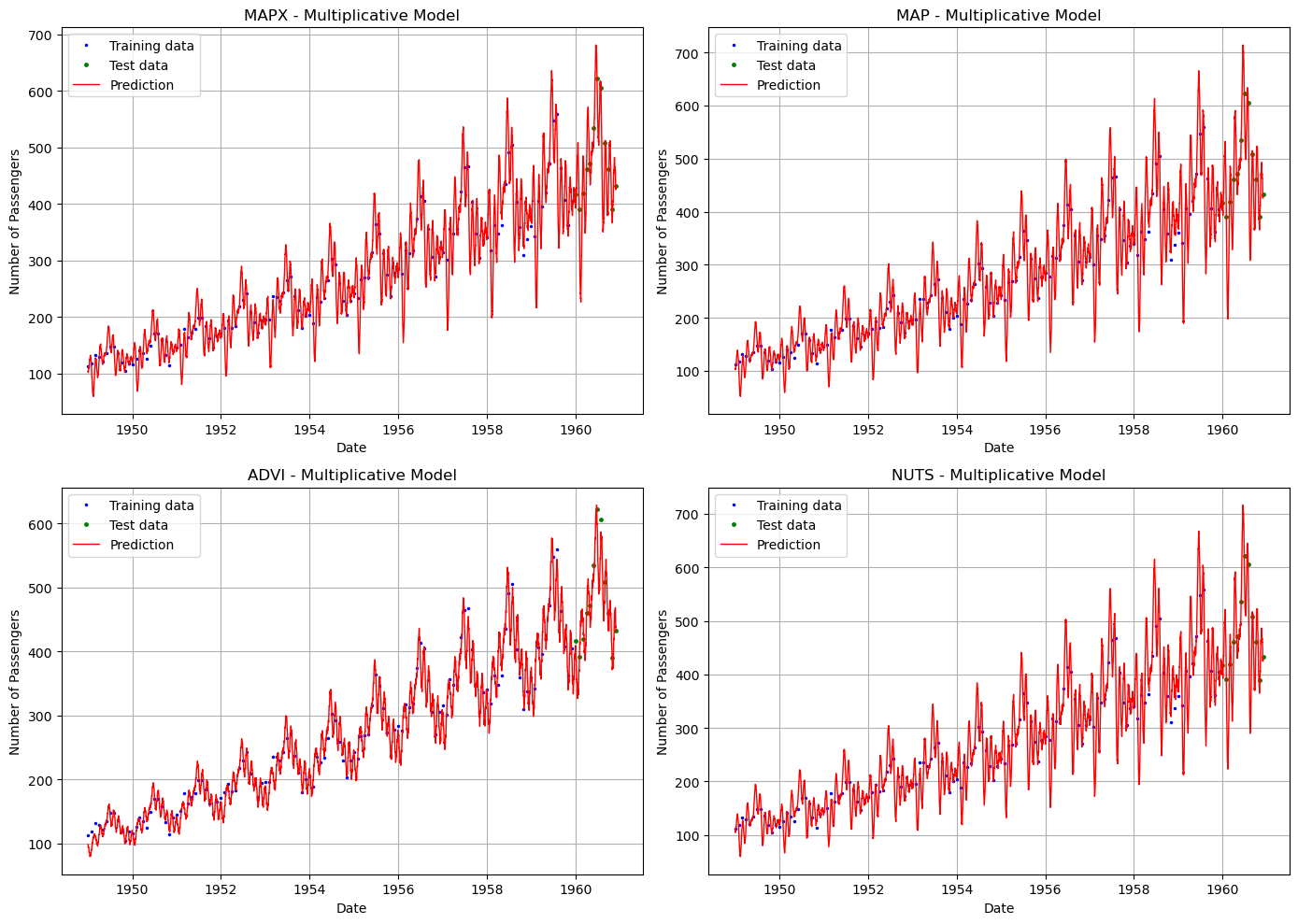
[14]:
# Visual comparison of predictions
fig, axes = plt.subplots(2, 2, figsize=(14, 10))
methods = [
("MAPX", future_mapx),
("MAP", future_map),
("ADVI", future_advi),
("NUTS", future_nuts),
]
for ax, (name, future) in zip(axes.flat, methods):
ax.plot(train["ds"], train["y"], "b.", label="Training data", markersize=3)
ax.plot(test["ds"], test["y"], "g.", label="Test data", markersize=5)
ax.plot(future["ds"], future["yhat_0"], "r-", label="Prediction", linewidth=1)
ax.set_title(f"{name} - Multiplicative Model")
ax.set_xlabel("Date")
ax.set_ylabel("Number of Passengers")
ax.legend()
ax.grid(True)
plt.tight_layout()
plt.show()
Summary: Choosing an Inference Method#
Method |
Speed |
Uncertainty |
Best For |
|---|---|---|---|
MAPX |
⚡⚡⚡ Fastest |
❌ None |
Quick prototyping, model selection, production with large data |
MAP |
⚡⚡ Fast |
❌ None |
Legacy compatibility |
ADVI |
⚡⚡ Fast |
✅ Approximate |
When you need uncertainty but MCMC is too slow |
NUTS |
⚡ Slow |
✅✅ Full |
Research, when accuracy matters most, small to medium data |
Transfer Learning Consideration#
When using vangja’s transfer learning capabilities (fitting short time series with priors from long time series), MCMC methods are particularly valuable because:
They capture the full posterior from the long time series
The posterior can be used directly as a prior for the short series
Uncertainty propagates correctly through the transfer process
What’s Next#
In Chapter 03, we explore vangja’s ability to fit multiple time series simultaneously using vectorized computations — a significant performance advantage over fitting each series one at a time.